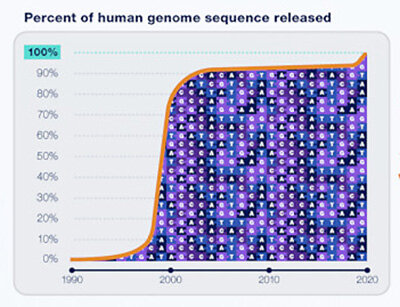
Scientists were a little ahead of their time when they pronounced the Human Genome Project complete two decades ago. With researchers throughout the world having access to the DNA sequences of most protein-coding genes in the human genome, a watershed moment had been achieved. Despite these advancements, 8% of the human genome remains unsequenced and unstudied after 20 years. The about 151 million base pairs of sequence data dispersed throughout the genome, dubbed “junk DNA” by some as having no evident function, remained a mystery.
In a report published in Science, a big international team led by Adam Phillippy at the National Institutes of Health has uncovered the remaining 8% of the human genome. More than just trash can be found in these long-missing parts of our genome. The latest findings reveal enigmatic regions of noncoding DNA that do not generate protein but play critical roles in many cellular functions and maybe at the root of diseases like cancer in which cell growth is out of control.
“You would think that, with 92 percent of the genome completed long ago, another eight percent wouldn’t contribute much,” says Rockefeller’s Erich D. Jarvis, a co-author on the study who helped develop a number of techniques central to unlocking the final pieces of the human genome. “But from that missing eight percent, we’re now gaining an entirely new understanding of how cells divide, allowing us to study a number of diseases we had not been able to get at before.”
On the shoulders of the HGP
The Human Genome Project essentially gave us the keys to euchromatin, the majority of the human genome that is densely packed with genes and busily producing RNA that would be translated into protein later. A maze of tightly wrapped, repeating heterochromatin—a smaller section of the genome that does not make protein—was left intact.
Scientists had solid reasons to put heterochromatin on the back burner at first. More genes were found in the euchromatic areas, which were also easier to the sequence. The genomics technologies of the time found euchromatic DNA easier to parse than its repeating, heterochromatic counterpart, much as a jigsaw with distinct parts is easier to put together than one with similar ones.
As a result, geneticists have a significant gap in their understanding of what drives some fundamental biological activities. In the human reference genome, the heterochromatic sequences behind centromeres, which reside at the cruxes of chromosomes and control cell division, were all annotated with long stretches of N for “unknown base.” The sequences of chromosomes 13, 14, 15, 21, and 22’s short arms were also removed. Jarvis continues, “Not even the entire euchromatic genome was sequenced adequately.” “Errors like false duplications had to be corrected.”
Then, about ten years ago, scientists began developing new techniques for producing longer sequence reads that filled in gaps in the genomes of humans and other species. One such initiative is the Vertebrate Genomes Project, helmed by Jarvis, which recently produced the first near error-free and near-complete reference genomes for 25 animals. “That study was part of an international effort to develop new tools that produce the highest-quality gene assemblies,” he says. “Compared to the methods that were used twenty years ago, modern genomics has high-fidelity long reads that are 99.9 percent accurate, better genome assembly tools, and more powerful algorithms that are better at distinguishing similar-looking puzzle pieces from one another.”
With updated tools and renewed resolve, Jarvis and other scientists were able to help finish what the Human Genome Project started and describe, at long last, a truly complete human genome—its euchromatic regions revised, and its heterochromatic regions on full display.
“It’s a big deal,” Jarvis says. “Every single base pair of a human genome is now complete.”
Meeting Merfin
The flagship Science study was led by the Telomere-to-Telomere (T2T) Consortium, a group of researchers at various academic institutions and NIH. The Jarvis lab’s contribution, published in Nature Methods, involved providing tools to help T2T refine messy genome sequences to produce error-free sequences.
One of these tools is Merfin, which they used to clean up some of the most difficult sequences in the human genome. “Genomes that we generate in the lab can have many errors in them,” says Giulio Formenti, a postdoc in Jarvis’ lab who developed Merfin. “If even just one or a few base pairs are wrong, that can have big consequences for the overall accuracy of the genomic sequence.” Merfin makes it possible to test the accuracy of a sequence, sensing code that may be out of place and automatically correcting mistakes. Because the technologies that generate modern sequences are more accurate, Merfin is reserved for only the trickiest cases.
“Stretches of identical base pairs, such as AAA, are hard for existing technology to assess,” Formenti says. “There are often errors in those sequences, even now. Merfin corrects them.”
Jarvis and Formenti hope that their contribution will not only help tie a bow on the Human Genome Project, but also inform research into diseases linked to the heterochromatic genome—chief among them cancer, which is associated with centromere abnormalities. Cancer cells divide wildly when certain heterochromatic centromere genes are overexpressed, and a complete understanding of the centromere genome may open the door to novel therapies.
“We are finally digging into what we once called junk DNA, because we could not understand it or look at it accurately,” Formenti says. “We now know that many diseases are linked to structural repeats in the centromere and, now that these sequences are no longer missing from the human reference genome, we can begin to map the origins of these diseases.”
Other co-authors in the Merfin study are: Arang Rhie, Brian P. Walenz, Françoise Thibaud-Nissen, Kishwar Shafin, Sergey Koren, Eugene W. Myers, and Adam M. Phillippy.
Source: Rockefeller University

